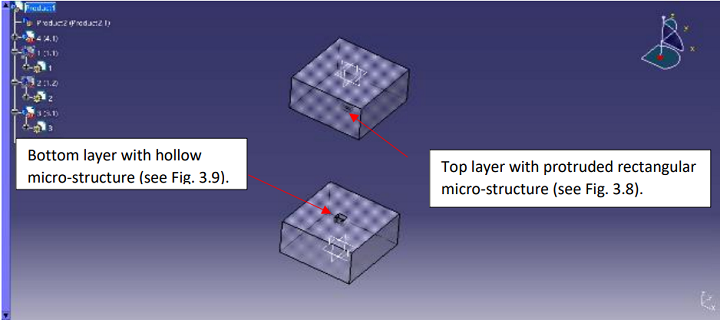
Polyjet 3D Printing Used to Make PDMS Molds for Sliding Trapper Microfluidics Design Concept
Microfluidics, which often crosses paths these days with 3D printing, deals with manipulating and controlling the flow of fluids in tiny channels, often on the sub-millimeter scale. Many microfluidic devices have been developed to assist in cell analysis, which is very beneficial for the medical field. For his University of Saskatchewan thesis paper, titled “On the Sliding Principle of Micro-Fluidic Devices for a Potential Use in Sorting Cells of Different Sizes with One Device,” Annal Arumugam Arthanari Arumugam focused on a new microfluidics design concept known as the sliding principle.
In the paper, Arumugam explains that while there are many microfluidic devices that can capture, isolate, position, and sort single cells, most can only work with cells of the same size. Tunable microfluidic devices can be used to capture and sort single cells sized 20 to 30 µm, but many applications have a desired size range of 2 μm to 100 μm, or even more.
“This thesis first conducted an analysis of different working principles of devices for capturing and sorting single cells, attempting a solution to the problem,” the thesis states. “As a result, this thesis proposed a novel principle for devices to perform sorting single cells with the cell size ranging from 2 μm to 100 μm, and this principle is named “sliding principle”. To prove the sliding principle to work, a device that contains a micro-trapper or well based on this principle was designed and fabricated using soft lithography with the mold fabricated with a 3D printing technology. The experiment conducted with the microscopy (resolution: 1-3 micron) and motion stage (resolution: 1 micron), which shows that the device can adjust the size of the well trapper, ranging from 0 to 1000 µm and covering the desired cell size range (i.e., 2 μm to 100 μm). According to the current literature on the mechanical approach to capture and sort single cells of different sizes with one device, the device built based on the sliding principle should potentially be applicable to capturing and sorting single cells of different sizes with one device.”
The device’s overall function requirement (FR) is to be able to capture cells of different sizes, ranging from 2 µ to 100 µ, with the resolution of 2-5 µm. Sub-function requirements included:
- Forming a sliding pair so the trapper changes the size from the sliding movement
- Executing a sliding trapper
- Pumping cell fluids to flow over the trapper
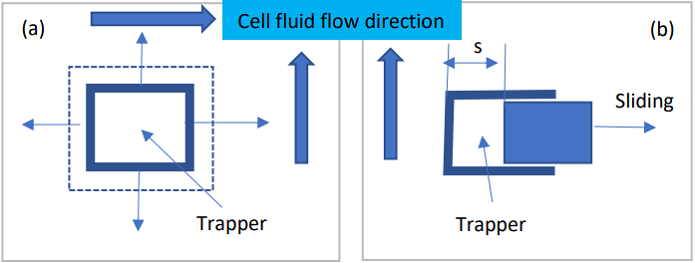
The sliding principle of the adaptable trapper. (a) The trapper is a square with four
edges that can be slid. (b) The one edge is sliding to change the size of the trappers.
The device that the cells contact had to be made from biocompatible material, with maximal stress in the cell at less than 4.5 Pa, and the adjusting range of the sliding less than 1000 µm. Arumugam considered two different design options for his sliding trapper, but the first did not work out, as the contact surface of two blocks may not have been flat enough to allow for smooth sliding between blocks and leakage was possible. So he instead focused on the second option.
“In this design, there are two sheets (top and bottom),” Arumugam explained. “On each sheet, there are a few wells (however, in this thesis only one well is considered without loss of generality), which are in the form of a square. Specifically, on the top sheet, the square is convex with a protruded part, and in the bottom sheet, the square is concave. After the two sheets are assembled (the top on the bottom), they form a system…”
A guide, rack, and top and bottom blocks, with embedded sheets made out of PDMS, make up the mechanism; a single axial stage with the motion resolution of ~3 um, made out of full cure 835 Vero white plus material, helped to drive the top block. Arumugam used Polyjet 3D printing to make a mold for the PDMS parts.
In testing the design, the device was measured to see if it met “the design specification in the geometry and topology of the device,” and the sliding operation was also measured, in order to “examine the change of the well.”
While the measurements were satisfactory for the PDMS sheets, and showed that the sliding principle concept is indeed valid, they were eroded a little on the sides, which makes channel spacing less accurate; the reason for this damage was due to the sticky PDMS not peeling cleanly away from the mold during curing.
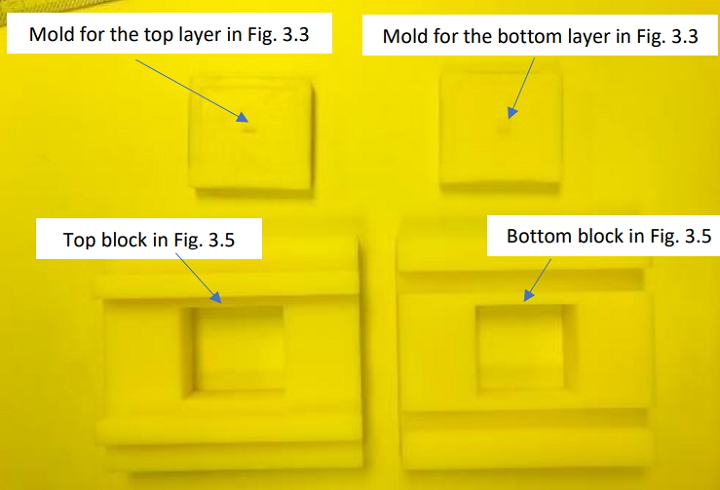
The 3D printed mold and sliding assembly for PDMS, which was made at the university’s engineering workshop.
Arumugam explained, “In the experiment, in the first few attempts, the PDMS did not solidify properly, and the PDMS sheet (molded part) was stick to the mold and damaged during the peel-off process. To address this problem, the 3D printed mold was prebaked in the oven for 85°C for 4 hrs and subsequent curing of PDMS sheet was performed. However, the problem has not completely gone away. This problem is responsible for inaccuracy (~2um) of the molded part in terms of dimension as well as surface damage. That resolution is partially due to the size of the channel, which is 1 mm. The size of the channel affects the focus of the microscopy, and subsequently the number of pixels that can be covered by the view, and thus the resolution in particular the length per pixel (8.547um now). Suppose that the maximal channel size is 100 µm. Then the measurement resolution will be 0.855 um.”
The author recommends some future works to help advance microfluidic device technology, such as optimizing the fabrication of PDMS channels and further modification of his design.
Discuss this research and other 3D printing topics at 3DPrintBoard.com or share your thoughts below.
Subscribe to Our Email Newsletter
Stay up-to-date on all the latest news from the 3D printing industry and receive information and offers from third party vendors.
Print Services
Upload your 3D Models and get them printed quickly and efficiently.
You May Also Like
Srini Kaza Discusses Strategically Scaling Align’s “Smile-Changing” 3D Printed Aligners
Align Technology‘s Invisalign is a revolutionary method to get you the smile you want through 3D printing. It is also a hugely popular process to go through, a $4 billion...
Analysis: Additive Manufacturing Strategies Spotlights Vertical Value Creation
A slowdown in the industry and particularly a tightening of access to capital following the additive manufacturing (AM) industry’s peak special purpose acquisition company (SPAC) phase in early 2021 is...
Department of Defense Awards PROTECT3D $1.3M for Custom 3D Printed Braces
Custom prosthetics company PROTECT3D has secured a $1.3M Department of Defense (DoD) Small Business Innovation Research contract to develop a platform for creating custom braces for soldiers. The PROTECT3D Platform...
AMS 2025 Highlights Big Changes in 3D Printing with Pivotal Speakers and Panels
2023 was filled with excitement around the potential mergers being pursued by industry stalwart Stratasys (Nasdaq: SSYS), leading to some of the most insightful conversations imaginable at Additive Manufacturing Strategies...